Two major advances in ocular health research – a newly awarded grant and a published database – have recently come out of the Medical College of Georgia at Augusta University, paving the way for significant improvements in patient diagnosis and treatment.
A grant to revolutionize dry eye disease care
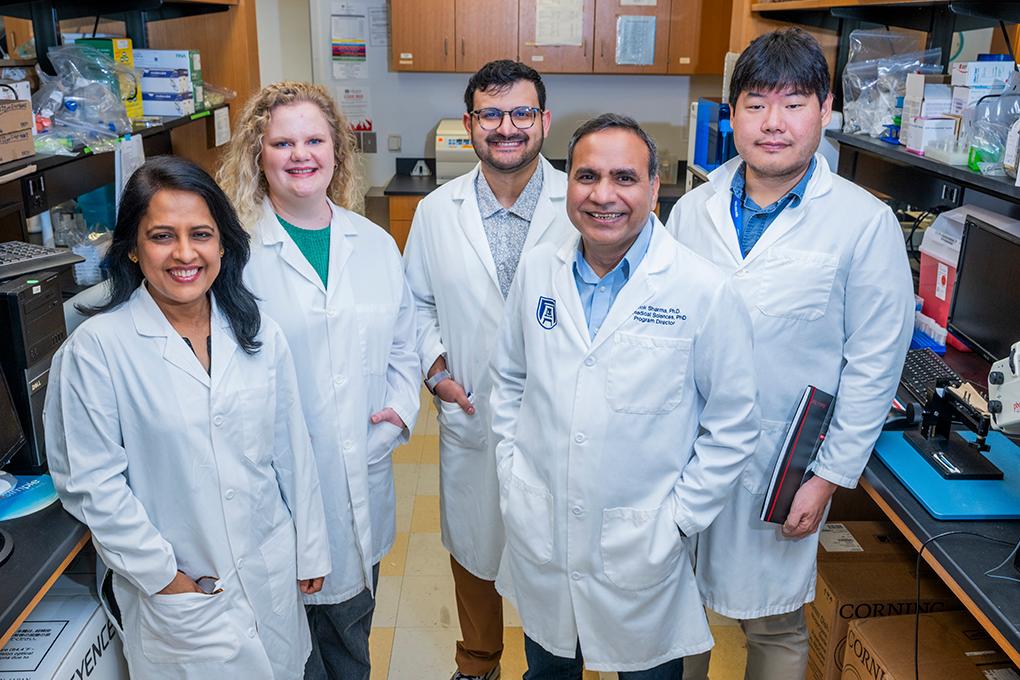
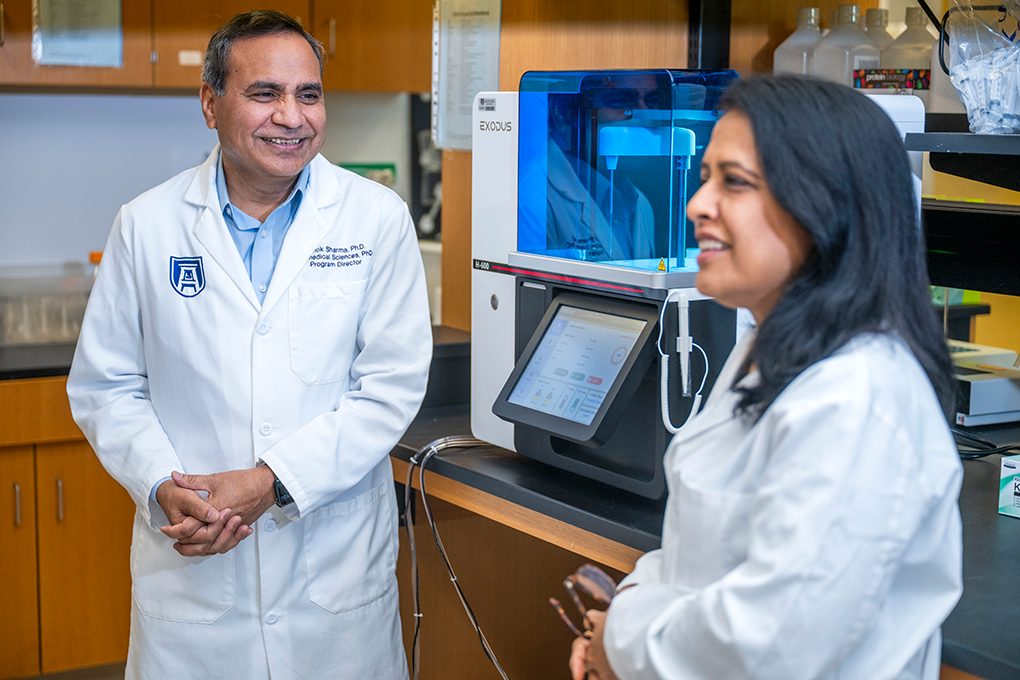
The nearly $1.5 million NIH R01 grant is the most recent of the two. It was awarded to husband and wife co-principal investigators Ashok Sharma, PhD, and Shruti Sharma, PhD, with Amy Estes, MD, and Vishal Jhanji, MD serving as co-investigators. This funding will allow them to study the mechanisms of dry eye disease (DED), aiming to provide more specialized treatments for those who suffer from the condition.
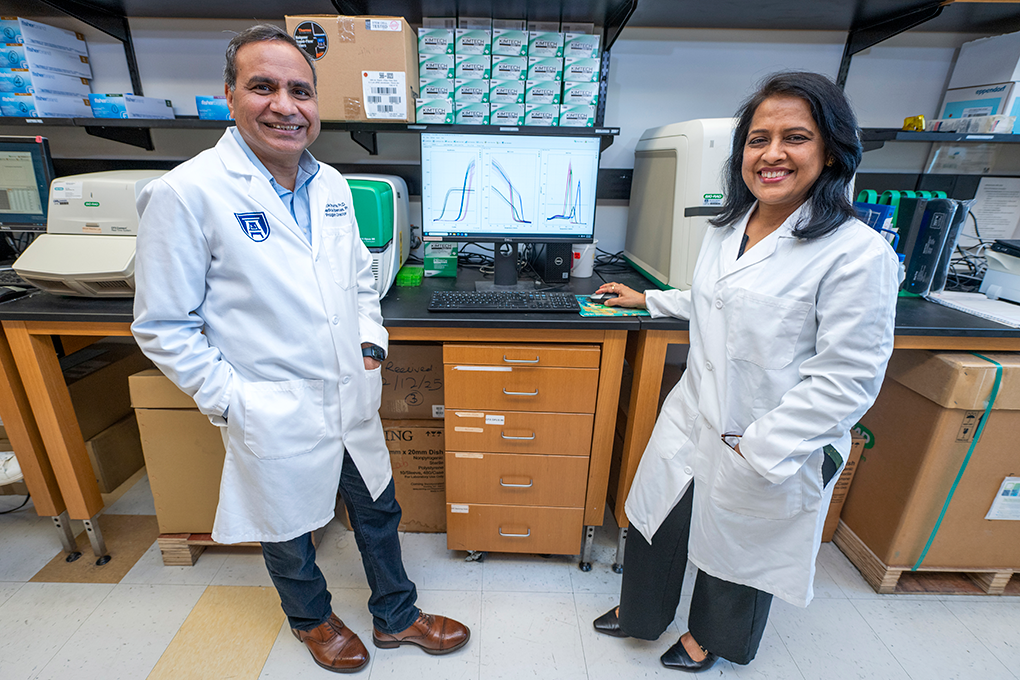

The study, titled “Molecular subtyping of dry eye disease: a machine learning approach using tear fluid proteins,” builds upon previous tear fluid research conducted by Ashok Sharma, professor and associate director of the MCG Center for Biotechnology and Genomic Medicine.
DED is a common condition affecting up to 30 million Americans in which there is inadequate lubrication of the eye due to low tear production, excessive evaporation or both. Symptoms include eye dryness, burning, irritation, blurred vision, redness, mucus, light sensitivity, a gritty sensation or, ironically, watery eyes.
Despite its prevalence, the disease remains challenging to diagnose and treat. There are many underlying causes that could result in DED, such as aqueous tear deficiency, abnormalities in the tear lipid layer and other systemic diseases. Signs and symptoms are often inconsistent and vary greatly, making it hard for clinicians to determine the best route for treatment.
Rather than relying solely on patients’ symptoms, the study aims to shift the diagnostic process toward examining the molecular composition of tears. The research team seeks to subtype DED patients based on their molecular profiles to allow clinicians to prescribe individualized medications based on causes, not symptoms, moving away from the current “one treatment fits all” approach.
“‘One drug fits all’ doesn’t work for dry eye disease because there are multiple factors that can lead to it. That’s why this is so important,” said Shruti Sharma, professor in the Department of Ophthalmology and the Center for Biotechnology and Genomic Medicine at MCG. “Right now, clinicians know there are different subtypes of dry eye disease based on the symptoms, yet they don’t have treatments for them. That’s why we think this study is a game changer.”
“For instance, if we know a patient’s dry eye disease is caused by inflammatory proteins, we can prescribe anti-inflammatory medications; if it’s related to autoimmune diseases, we can prescribe something else,” Ashok Sharma added.
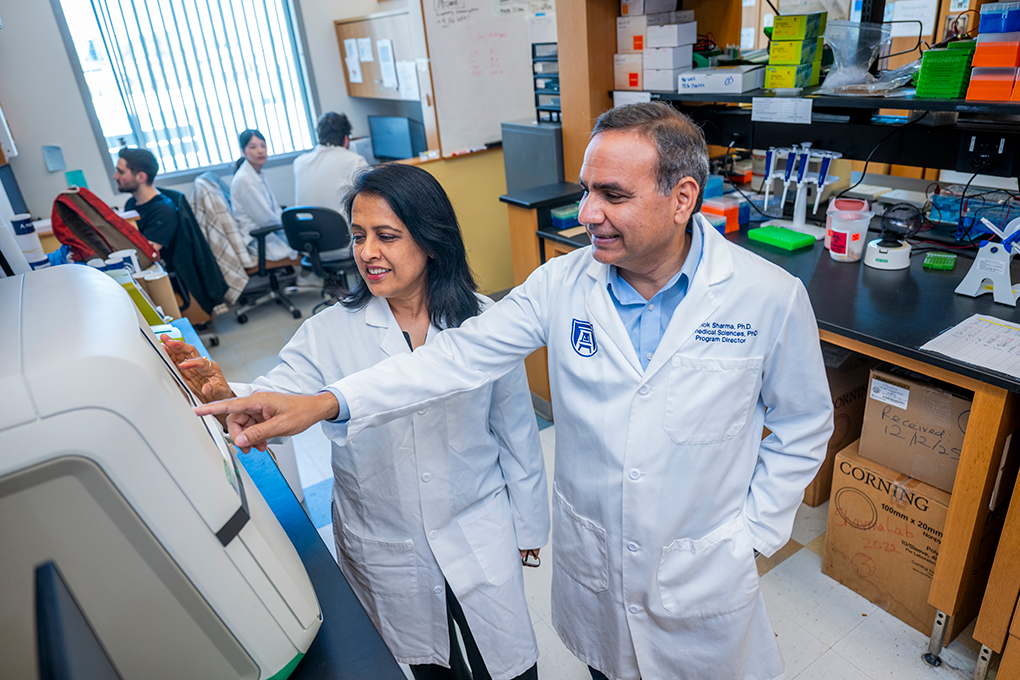

To subtype the patients in the study, the team will use mass spectrometers to generate molecular profiles for each of them based on their tear fluid. Computational approaches will then be used to identify patterns within those profiles and group patients with similar profiles together.
The project also aims to develop a point-of-care diagnostic tool based on multiplex immunoassays, which are techniques that simultaneously measure multiple analytes within a single small-volume sample. This approach will allow expedited, real-time classification of dry eye disease in clinical settings.
“Testing those proteins in the clinic will allow the study to move from research to bed side,” Ashok Sharma said.
Looking to the future, the team aspires to apply their tear fluid research and protein analysis workflow to other diseases with the goal of developing better treatment options. Previous studies show that changes in the molecular composition of tear fluid could be linked not only to DED and other ocular ailments, but also to diabetes, breast cancer and even neurodegenerative diseases like Alzheimer’s and Parkinson’s.
“The eye shares developmental and physiological connections with the central nervous system, which may enable tear fluid to reflect neurological conditions,” Shruti Sharma said. “That’s why it makes sense that tear fluid could be linked to brain-related diseases.”

A comprehensive tear fluid database
The dry eye disease study wouldn’t be possible without another recent major scientific accomplishment of Ashok Sharma’s: a tear fluid database.
Tear fluid is a not-often-thought-about liquid that our eyes rely on daily – it lubricates them, protects them from harmful bacteria, delivers oxygen and other nutrients to them and helps to heal surface-level injuries. But, Sharma and his lab team discovered that there is so much more that makes tear fluid special.
There are four main types of tears: emotional, reflex, closed-eye and basal, which is the type Sharma’s team is studying due to their constant presence on the surface of the eye.
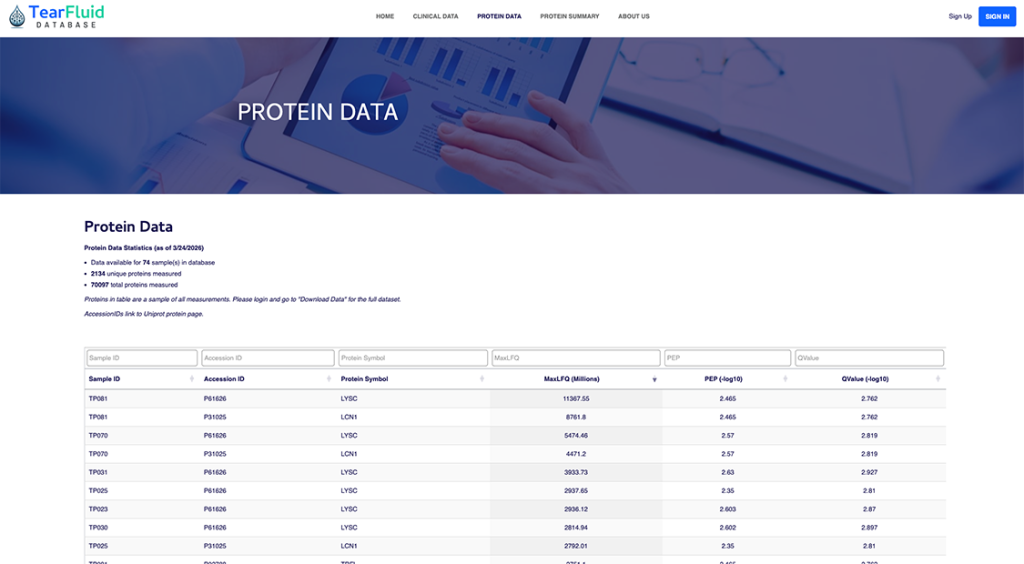
The basal tear fluid is a complex three-layered film containing more than a thousand different types of proteins, hundreds of microRNAs and other molecules. The team shared these results by developing a comprehensive tear fluid database and launching a companion website. They then published an article about their findings in the scientific journal Database titled, “Tear fluid database: a reference website for tear fluid proteomics.”
They also recently published an article in The Ocular Surface, “Comprehensive profiling of the human tear fluid miRNome using small RNA sequencing,” about their findings of different microRNA molecules present in tear fluid and whether they vary by criteria such as sex, age and race.
Sharma compares tear fluid analysis to getting routine blood work, noting that blood contains molecules and other biomarkers linked to certain diseases.
“Similarly, we want to implement tear work one day as well. So, a doctor will collect tears from your eyes and analyze them to see whether you are healthy or if you have a certain kind of abnormal condition,” he explained.

The difficult part of analyzing this type of tear fluid is collecting it. Because there is such a small amount on the eye’s surface – less than a drop – scientists have to collect it using a paper strip called a Schirmer strip.
“We collect the tears on this piece of paper, then put it in a tube, extract all the molecules out of it and analyze them,” Sharma explained. “We are in the era of technological development right now, and very high-quality mass spectrometers here at MCG can now detect thousands of proteins from less than a drop of tears absorbed within the paper strip.”
The team spent four years optimizing the extraction method for the proteins and molecular information in tear fluid before launching their database and website. From 74 human tear samples, the database lists all 2,134 proteins identified in the fluid thus far and their amounts. The most abundant is Lipocalin-1 and the second most abundant is Lysozyme C – both have anti-microbial properties that help protect the eyes.
Not only is the database comprehensive, it’s also user-friendly, allowing users to search for specific proteins in the tear fluid based on their protein symbols.
Michael F. Chiang, MD, the director of the National Eye Institute, recently gave a nod to Sharma’s work on X – exposing his research to broader audiences and putting more eyes on the work being done at MCG.

“This represents one of the first comprehensive efforts to develop a website and database dedicated to tear fluid proteins,” Ashok Sharma said. “How do these proteins in tear fluid change in a disease condition? If they are changing, are they increasing or decreasing? Can we try to use them as biomarkers or therapeutic targets?”
He also noted that some companies are currently trying to develop wearable devices such as contact lenses that can continuously monitor the biomarkers in tear fluid.
He, Shruti and their lab members are looking forward to applying their groundbreaking tear fluid research to other debilitating diseases in hopes to improve patients’ quality of life.
“The ultimate goal is to analyze and classify tear samples so patients can be grouped based on the underlying subtype of disease and treated with the most appropriate therapy,” he said. “If successful, this approach could be transformative by enabling personalized treatment rather than relying on trial and error.”




 Augusta University
Augusta University