Six novel chromosomal regions identified by scientists leading a large, prospective study of children at risk for type 1 diabetes will enable the discovery of more genes that cause the disease and more targets for treating or even preventing it.
The TEDDY study’s international research team has identified the new gene regions in young people who have already developed type 1 diabetes or who have started making antibodies against their insulin-producing cells, often a precursor state to the full-blown disease that leads to a lifetime of insulin therapy.
Their analysis of 5,806 individuals published in the Journal of Autoimmunity also confirmed three regions already associated with one of those related conditions.
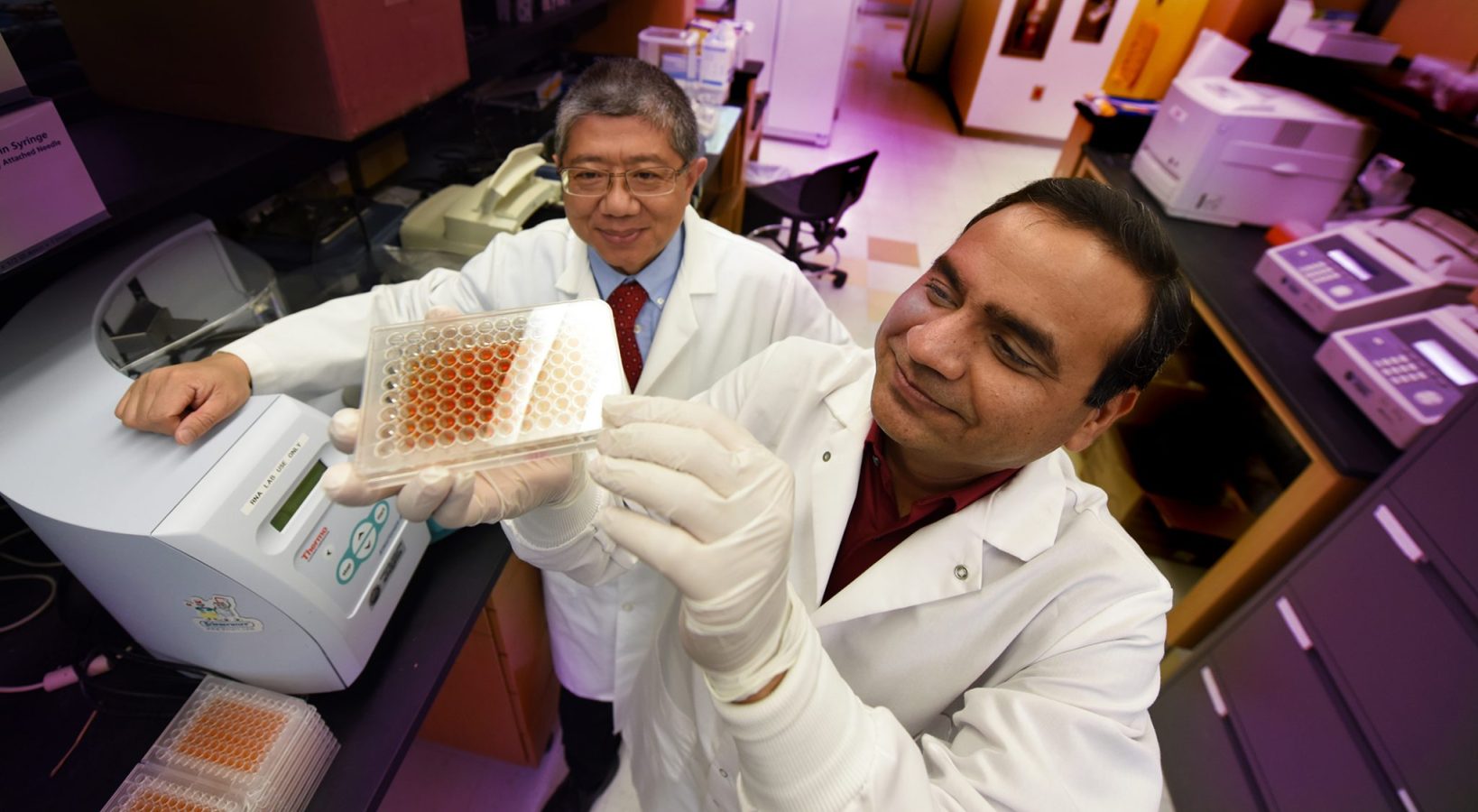
“We want to build a more precise profile of who will get this disease and when,” says Dr. Jin-Xiong She, director of the Center for Biotechnology and Genomic Medicine at the Medical College of Georgia at Augusta University, principal investigator of TEDDY’s Georgia/Florida site and the study’s corresponding author.
In keeping with their theory that two subtypes of type 1 diabetes will become clear from longitudinal studies of those at risk, the international TEDDY team also found different chromosomal regions were associated with which autoantibody shows up first in a patient, a sign his immune system is turning on his pancreas.
They looked at two top autoantibodies: one directly against insulin, called IAA, and one called GADA, against the enzyme glutamate decarboxylase, which regulates the insulin-producing beta cells in the pancreas. About 90 percent of patients with type 1 diabetes have one or the other autoantibody first and many eventually end up with both, She says. The second autoantibody may surface in a few days or even years later.
“There is mounting evidence that we have at least two major subtypes of type 1 diabetes, based on the autoantibodies children have. Now we have found a genetic basis that supports that,” says She, Georgia Research Alliance Eminent Scholar in Genomic Medicine.
TEDDY – The Environmental Determinants of Diabetes in the Young – is an international initiative following almost 9,000 children for 15 years in a strategic and rare opportunity to watch how genetics and environmental factors collide to cause disease, She says. An original goal of TEDDY was to better determine which genetic variations correlate with progression or lack of progression to type 1 diabetes.
For this particular pursuit, they focused on the 5,806 Caucasian TEDDY participants, because of genetic differences in different ethnic groups. They also focused on non-HLA genes, says Dr. Ashok Sharma, MCG bioinformatics expert and the study’s first author.
Most genes known to be associated with type 1 diabetes – including those currently considered the top two high-risk genes, which are the ones TEDDY screens for – are classified as human leukocyte antigen, or HLA genes. It’s a logical association since HLA genes regulate our immune system, says Sharma.
But in their comprehensive effort to better identify children at highest risk of disease – and ideally one day intervene – this particular search focused on non-HLA genes. “By study design we were looking for them,” Sharma says.
“With HLA genes you can achieve a certain level of accuracy in identifying high-risk individuals,” says She. “But if we can add additional genes into the screening, we can refine the prediction of the disease, we can increase the accuracy, we can probably even identify higher percentages of at-risk individuals.”
“It’s not monogenic, there are many genes involved,” Sharma says of type 1 diabetes, a condition that affects 1 in 300 people in the United States by age 18, according to the National Institutes of Health.
Which of those genes are involved also varies by individual. Sharma and She note the reality that not all patients with the high-risk genes even get the disease, although they still don’t know why.
One of the many points they hope TEDDY clarifies is if or how these genetics along with environmental factors – like childhood infections or even what children eat – conspire to cause actual disease. Genetic factors are comparatively easier to identify, and should also help identify environmental factors, they say.
For this study, the scientists also started with 176,586 SNPs, or single nucleotide polymorphisms. A nucleotide is a basic building block of our genetic information. In the case of DNA, those are A, C, T and G, chemical bases that can be arranged in seemingly endless potential orders. SNPs are genetic variations – one letter replaced by another – that scientists are using increasingly to figure out which occur more or less often in people with a certain condition or disease.
The SNPs examined by TEDDY scientists were already associated with other autoimmune diseases like rheumatoid arthritis or celiac disease, but not type 1 diabetes, Sharma says. They determined which of these SNPs are different in TEDDY participants who had developed type 1 diabetes versus those who had Islet cell autoantibodies versus those who still had neither.
Previous research has shown that the same genes are not always associated with IA and actual type 1 diabetes.
In fact, while Islet cell autoantibodies, or IA, are considered a red flag for type 1 diabetes, not every child with IA will progress to full-blown disease, She says, although multiple autoantibodies definitely increase that risk. In fact, different genes may play a role in IA development while others play a role in disease progression, the TEDDY scientists write.
While the SNPs themselves may or may not have a direct functional consequence, they are in close proximity to a gene that does, says She. “It’s a marker,” he says, that will enable the scientists to more efficiently and effectively identify causative genes.
“We are using SNPs but not as an endpoint,” Sharma notes. “We want to find out the genes which are there,” a focal point for the work now underway.
Key to the work already complete, is how closely and long TEDDY participants are followed. Normally gene identification starts with what is called a case-control design, in which genetic variations are compared between patients with a condition and healthy individuals, to look for differences that may contribute to the disease.
While standard cross-sectional, or case-control studies haven’t shown a huge impact from non-HLA genes, they only provide a “snapshot’ of what is happening with both patients and controls alike, the scientists say. With the prospective and longitudinal TEDDY, the scientists are literally watching the disease occur – or not – in high-risk young people.
In fact, this is the first major study regarding gene identification for any disease that uses this sort of longitudinal information, She says.
As with many things, timing is everything, and the TEDDY perspective sharpens the search, particularly for important non-HLA genes by adding the “time to disease” perspective, She says.
Co-authors include leading diabetes researchers in Georgia, Florida, Colorado, Washington, Finland, Sweden and Germany. In addition to MCG, lead institutions include Morsani College of Medicine at the University of South Florida; Pacific Northwest Research Institute in Seattle; Sweden’s Lund University; the University of Colorado; Germany’s Helmholtz Zentrum München and Klinikum rechts der Isar, Technische Universitӓ München and Forschergruppe Diabetes e.V.; and Finland’s University of Turku and Turku University Hospital. She has an appointment in the MCG Department of Obstetrics and Gynecology and Sharma is in the Department of Population Health Sciences.
TEDDY has been funded since 2003 by the NIH grants U01 DK63829, U01 DK63861, U01 DK63821, U01 DK63865, U01 DK63863, U01 DK63836, U01 DK63790, UC4 DK63829, UC4 DK63861, UC4 DK63821, UC4 DK63865, UC4 DK63863, UC4 DK63836, UC4 DK95300, UC4 DK100238, UC4 DK106955, UC4 DK112243, UC4 DK117483, and Contract No. HHSN267200700014C.
 Augusta University
Augusta University